Difference between revisions of "FlyBase:Tools Overview"
| (292 intermediate revisions by 13 users not shown) | |||
| Line 1: | Line 1: | ||
=General Search Help and Tips= | =General Search Help and Tips= | ||
| − | |||
| − | |||
FlyBase can be searched for genes, alleles, aberrations and other genetic objects, phenotypes, sequences, stocks, images and movies, controlled terms, and Drosophila researchers using the tools available from the 'Tools' drop-down menu in the Navigation bar. In addition to the Navigation bar, which can be accessed from any FlyBase page, the homepage also has direct links to the most commonly used tools. | FlyBase can be searched for genes, alleles, aberrations and other genetic objects, phenotypes, sequences, stocks, images and movies, controlled terms, and Drosophila researchers using the tools available from the 'Tools' drop-down menu in the Navigation bar. In addition to the Navigation bar, which can be accessed from any FlyBase page, the homepage also has direct links to the most commonly used tools. | ||
| − | Below are | + | Below are brief descriptions of each of the tools, which have been split into five main sections: |
*[[FlyBase:Tools Overview#Overview of Search Strategies|Overview of Search Strategies]] (for example, how to search for expression data) | *[[FlyBase:Tools Overview#Overview of Search Strategies|Overview of Search Strategies]] (for example, how to search for expression data) | ||
*[[FlyBase:Tools Overview#Main Query Tools|Main Query Tools]] (Jump to Gene, QueryBuilder, etc.) | *[[FlyBase:Tools Overview#Main Query Tools|Main Query Tools]] (Jump to Gene, QueryBuilder, etc.) | ||
*[[FlyBase:Tools Overview#Query Results Analysis Tools|Query Results Analysis Tools]] (Hit list refinement, Batch Download) | *[[FlyBase:Tools Overview#Query Results Analysis Tools|Query Results Analysis Tools]] (Hit list refinement, Batch Download) | ||
| − | *[[FlyBase:Tools Overview#Genomic Search Tools and Browsers|Genomic Search Tools and Browsers]] ( | + | *[[FlyBase:Tools Overview#Genomic Search Tools and Browsers|Genomic Search Tools and Browsers]] (JBrowse, BLAST etc.) |
| − | *[[FlyBase:Tools Overview#Other Tools|Other Tools]] (Interactions Browser, | + | *[[FlyBase:Tools Overview#Other Tools|Other Tools]] (Interactions Browser, Fast Track Your Paper etc.) |
| + | |||
| + | Links to '''Full documentation''' for FlyBase tools can be found in the [[FlyBase:FlyBase_Help_Index#Tools_and_Downloads_Documentation | Tools section of the FlyBase Help Index]]. | ||
=Overview of Search Strategies= | =Overview of Search Strategies= | ||
| Line 17: | Line 17: | ||
==Searching 12 species== | ==Searching 12 species== | ||
| − | + | Please note -- Starting in 2018, FlyBase will reflect updated gene models annotated by the NCBI gnomon pipeline for four species only: D. simulans, D. ananassae, D. pseudoobscura, and D. virilis. Thus, existing gene records for the other seven AAA species may go stale and newly annotated genes will not be included. (D. melanogaster gene models are updated via a separate pipeline.) This section will be updated as these changes occur. | |
| − | + | Individual gene reports for genes from the 12 originally sequenced Drosophila genomes are available in FlyBase. There are four main ways in which these data can be browsed and queried in FlyBase: | |
| − | *[http:// | + | *[http://flybase.org/downloads/bulkdata Precomputed files] |
| − | *[http://{{ | + | *[http://{{flybaseorg}}/blast/ BLAST] |
*Gene Report Pages | *Gene Report Pages | ||
| Line 29: | Line 29: | ||
For those interested in genome-wide analyses, bioinformatics and comparative genomics, there are a selection of pre-computed files available for download from our precomputed files page (in the Genomes:Annotation and Sequence section, for example), found in the 'Files' menu. | For those interested in genome-wide analyses, bioinformatics and comparative genomics, there are a selection of pre-computed files available for download from our precomputed files page (in the Genomes:Annotation and Sequence section, for example), found in the 'Files' menu. | ||
| − | For those with an interest in a specific gene/protein/region across the different species, there are a number of ways to query the data. Our [http://{{ | + | For those with an interest in a specific gene/protein/region across the different species, there are a number of ways to query the data. Our [http://{{flybaseorg}}/blast/ BLAST server] allows querying of numerous sequenced insect genomes, either individually, as a subset, or all together. |
| − | ==Aberrations - deficiencies, inversions, translocations== | + | ==Aberrations - deficiencies, dupications, inversions, translocations== |
| − | One of the problems in a field of the size and complexity of Drosophila genetics is the use of nomenclature. This can lead to a number of names being given to the same object, and to the valid FlyBase name or symbol of an object being quite confusing or indeed not in common lab parlance. Aberration naming is no exception. The simplest ways to search for an aberration are either using [http://{{ | + | One of the problems in a field of the size and complexity of Drosophila genetics is the use of nomenclature. This can lead to a number of names being given to the same object, and to the valid FlyBase name or symbol of an object being quite confusing or indeed not in common lab parlance. Aberration naming is no exception. The simplest ways to search for an aberration are either using [http://{{flybaseorg}}/cytosearch CytoSearch], when you want to find an aberration that removes a particular gene or uncovers a cytological band, or using [http://{{flybaseorg}}/ QuickSearch] (use the 'Data Class' tab and select 'aberration' as the data class). Remember to use wildcards (i.e. *) to allow for slight differences in naming. FlyBase records all mentions of an aberration, so if an aberration is given a particular symbol in a paper, this name will be recorded as a synonym of the FlyBase 'valid' symbol (see the [http://{{flybaseorg}}/wiki/FlyBase:Nomenclature nomenclature] document for more details). Alternatively, you can browse the molecularly localized aberrations for each chromosome by scanning [http://{{flybaseorg}}/cgi-bin/gbrowse2/dmel/ JBrowse] after selecting all "Aberrations" tracks. |
| − | ==Cytologically | + | ==Cytologically Mapped Features== |
| − | When looking for cytology, you have a choice of a number of tools on FlyBase, including [http://{{ | + | When looking for cytology, you have a choice of a number of tools on FlyBase, including [http://{{flybaseorg}}/cgi-bin/qb.pl QueryBuilder]. The easiest tools to use however, are [http://{{flybaseorg}}/cytosearch CytoSearch] or [http://{{flybaseorg}}/cgi-bin/gbrowse2/dmel/ JBrowse]. JBrowse is especially useful when looking for molecularly mapped sequences, insertions, or probes. CytoSearch comes into its own when searching for cytologically defined features, such as cytologically-mapped genes or deficiencies, that haven't been molecularly mapped to the sequence. Of course, as with many aspects of research, complimentary methods should be used. Therefore, we recommend you use both JBrowse and CytoSearch to analyse cytology. |
| + | |||
| + | A description of how FlyBase computes the cytological location for features that have been mapped to the genome can be found in [[FlyBase:Computed cytological data | Computed cytological data]]. Illustrations or electron micrographs of ''D. melanogaster'' polytene chromosomes as well as a cytogenetic-genetic-sequence location correspondence table can be found at [[FlyBase:Maps | ''D. melanogaster'' Chromosome Maps]]. | ||
==Expression Data== | ==Expression Data== | ||
| Line 43: | Line 45: | ||
'''Browsing Expression Data''' | '''Browsing Expression Data''' | ||
| − | Expression patterns are captured by FlyBase curators for transcripts, proteins, and "reporters" (i.e. enhancer trap insertions and reporter constructs). Information about transcript and protein expression patterns can be found on gene reports (e.g. the [http://{{ | + | Expression patterns are captured by FlyBase curators for transcripts, proteins, and "reporters" (i.e. enhancer trap insertions and reporter constructs). Information about transcript and protein expression patterns can be found on gene reports (e.g. the [http://{{flybaseorg}}/reports/FBgn0260400.html elav] gene), data for reporter constructs can be found on recombinant construct reports (e.g. [http://{{flybaseorg}}/reports/FBtp0011922.html P{elav-lacZ.H}]) and associated allele reports (e.g. [http://{{flybaseorg}}/reports/FBal0047660.html Ecol\lacZ<sup>elav.PH</sup>]), and data for enhancer or protein traps can be found on insertion reports (e.g. [http://{{flybaseorg}}/reports/FBti0002575.html P{GawB}elav<sup>C155</sup>]) and associated allele reports (e.g. [http://{{flybaseorg}}/reports/FBal0047071.html Scer\GAL4<sup>elav-C155</sup>]). In all cases, expression data will be found in the "Expression Data" section of the report. For those constructs or insertions that reflect expression of a particular gene, data are also promoted to the corresponding gene report, in a subsection of "Expression Data" labeled "Expression Deduced from Reporters" (e.g. expression data for both [http://{{flybaseorg}}/reports/FBtp0011922.html P{elav-lacZ.H}] and [http://{{flybaseorg}}/reports/FBti0002575.html P{GawB}elav<sup>C155</sup>] are displayed on the [http://{{flybaseorg}}/reports/FBgn0260400.html elav] gene report). Subcellular Localization of protein is populated from [https://wiki.flybase.org/wiki/FlyBase:Gene_Report#Function Gene Ontology (GO) Cellular Component] curation of genes. |
| − | We | + | We cooperate with several other databases of expression data and either display a portion of their data within FlyBase (e.g. [http://www.flyexpress.net/ FlyExpress]) and/or link to their database (e.g. [http://fly-fish.ccbr.utoronto.ca/ Fly-FISH]). These types of data can be found in the "External Data & Images" subsection of the "Expression Data" section. Additionally, we maintain a set of links to [http://{{flybaseorg}}/wiki/FlyBase:Images Image Based Resources], including image databases, tools for image analysis, and tools for image visualization and annotation. |
| − | High throughput expression data | + | High throughput expression data from [http://flyatlas.org/atlas.cgi FlyAtlas] and [http://www.modencode.org/ modENCODE] can be found on Gene Reports in a subsection of "Expression Data" labeled "High-Throughput Expression Data". The modENCODE data can be visualized as a linear or log graph, or as a heatmap. The FlyAtlas section also includes a 'back-to-back' option, in which gene expression levels in larval tissues are juxtaposed with gene expression levels in the corresponding adult tissues. The graph displays can be scaled by gene maximum expressed, or by low, moderate, or high expression bin max. |
'''Searching for Expression Patterns''' | '''Searching for Expression Patterns''' | ||
| − | Expression data can be searched most easily and accurately by using [http://{{ | + | Expression data curated from literature can be searched most easily and accurately by using the QuickSearch [http://{{flybaseorg}}/wiki/FlyBase:QuickSearch#Expression_tab Expression] or [http://{{flybaseorg}}/wiki/FlyBase:QuickSearch#GAL4_etc_tab Gal4 etc] tabs (detailed help at links). The Expression tab allows searches for genes by temporal-spatial expression pattern, while the GAL4 etc allows searches for GAL4 and other binary drivers, and non-binary reporters. Another expression pattern search option is [http://{{flybaseorg}}/cgi-bin/qb.pl QueryBuilder], which supports multipart queries (e.g. generate a list of genes which have the GO term "transcription factor activity" and whose protein products are expressed in the central nervous system). However, if you're interested in all genes expressed in a bodypart, tissue, or developmental stage, you can find that using [http://{{flybaseorg}}/vocabularies Vocabularies]. For example, by entering the term "adult mushroom body" into Vocabularies, you can obtain a list of genes expressed in that tissue. |
'''Searching for High-Throughput Expression Patterns''' | '''Searching for High-Throughput Expression Patterns''' | ||
| − | RNA-Seq expression data can be searched to identify genes with specific expression characteristics using the | + | RNA-Seq expression data can be searched to identify genes with specific expression characteristics using the [http://{{flybaseorg}}/rnaseq/profile_search RNA-Seq Profile Search] tool. Genes that have expression patterns similar to a given gene can be found using the [http://{{flybaseorg}}/rnaseq/simsearch RNA-Seq Similarity Search] tool; this search option can also be launched from the relevant gene page. [http://{{flybaseorg}}/rnaseq/region RNA-Seq By Region] can be used to compare the RNA-Seq signal for a given region across samples, or to compare signal between two regions within a single sample. For additional information about these tools see [[FlyBase:RNA-Seq Overview | RNA-Seq Query Tools and Browsers]]. |
| − | ==Mutant | + | ==Mutant Phenotype Data== |
| − | Mutant phenotype data is associated with alleles in FlyBase, so you need to search allele data if you are interested in mutant phenotype. In addition to free text describing the phenotype, the alleles are indexed with [ | + | Mutant phenotype data is associated with alleles in FlyBase, so you need to search allele data if you are interested in mutant phenotype. In addition to free text describing the phenotype, the alleles are indexed with [[FlyBase:Controlled vocabularies used by FlyBase | controlled vocabulary]] (CV) terms, which makes it easier for you to search for a particular phenotype, e.g. searching for mutants that affect the wing. You can search with these CV terms using either [http://{{flybaseorg}}/vocabularies Vocabularies] or [http://{{flybaseorg}}/cgi-bin/qb.pl QueryBuilder]. |
You can find mutant alleles affecting the wing from all species using Vocabularies. If you enter the term "wing" into Vocabularies search page and then click on the "Alleles" button in the report page, you will obtain a list of mutant alleles that affect the wing. However, to search in a specific species, or to search for mutant phenotypes as part of a multipart query, QueryBuilder must be used. In this case, you should pick the "CV Hierarchy (GO/etc.)" dataset and then use the term picker to choose the body part, e.g. wing. In both cases, the default is to search both for alleles specifically labelled with the CV term, e.g. wing and also with child CV terms that are a subset of the term chosen, e.g. wing vein. If you want to restrict your search to just the precise term chosen, use QueryBuilder and select 'Retrieve records annotated with "This CV term only"' before you run the query. | You can find mutant alleles affecting the wing from all species using Vocabularies. If you enter the term "wing" into Vocabularies search page and then click on the "Alleles" button in the report page, you will obtain a list of mutant alleles that affect the wing. However, to search in a specific species, or to search for mutant phenotypes as part of a multipart query, QueryBuilder must be used. In this case, you should pick the "CV Hierarchy (GO/etc.)" dataset and then use the term picker to choose the body part, e.g. wing. In both cases, the default is to search both for alleles specifically labelled with the CV term, e.g. wing and also with child CV terms that are a subset of the term chosen, e.g. wing vein. If you want to restrict your search to just the precise term chosen, use QueryBuilder and select 'Retrieve records annotated with "This CV term only"' before you run the query. | ||
| + | |||
| + | Find a related '''video tutorial''' at [https://www.youtube.com/watch?v=hZgsDPypZvk Finding genes with similar phenotypes]. | ||
==References== | ==References== | ||
| − | FlyBase is an excellent source of Drosophila references. References can be searched in a number of ways. The easiest way is through [http://{{ | + | FlyBase is an excellent source of Drosophila references. References can be searched in a number of ways. The easiest way is through [http://{{flybaseorg}}/ QuickSearch], on our homepage. Choose the 'References' tab and fill in one or more of the search boxes. The field identity of each search box can be modified using the dropdown menus at the left. For more information, please go to the [http://flybase.org/wiki/FlyBase:QuickSearch QuickSearch Help Page]. |
| − | More refined reference searches can be performed using [http://{{ | + | More refined reference searches can be performed using [http://{{flybaseorg}}/cgi-bin/qb.pl QueryBuilder] (QB). Click on the box titled 'Query is empty.. Click here to start building' on the QB start page to being the search. At this stage the window will be displaying all the fields available to search for the 'Genes' dataset. Change the dataset to 'References'. Now the fields found in the reference reports are displayed. From here, you can search all the data found in the reference report, including pubmed ID, author, and type (e.g. review). Find complete instructions on the [http://{{flybaseorg}}/cgi-bin/qb.pl QueryBuilder] tool page. |
A popular way to search for references is to search for a (list of) objects (e.g. genes, GO terms) and then to use the 'Show related' toggle on the hits page to change the hit list to the related references. The 'Results Analysis/Refinement' button, found on the hit list page, can be used to analyse the distribution of the references over year, journal, author, and type of publication (e.g. review, paper, abstract). | A popular way to search for references is to search for a (list of) objects (e.g. genes, GO terms) and then to use the 'Show related' toggle on the hits page to change the hit list to the related references. The 'Results Analysis/Refinement' button, found on the hit list page, can be used to analyse the distribution of the references over year, journal, author, and type of publication (e.g. review, paper, abstract). | ||
| Line 73: | Line 77: | ||
==Stocks== | ==Stocks== | ||
| − | One of the easiest ways to search for a stock in FlyBase is to use [http://{{ | + | One of the easiest ways to search for a stock in FlyBase is to use [http://{{flybaseorg}}/ QuickSearch]. Simply change the data class to 'stocks', type in the feature of interest (e.g. a gene symbol, allele symbol), and search. A further way to identify stocks is through the hit list produced after a search. At the top of the hit list there is a toggle allowing you to 'Show related' stocks. Stocks can also be found for individual alleles by clicking on the Stocks matryoska on the allele report page. |
=Main Query Tools= | =Main Query Tools= | ||
| − | ==Jump to Gene== | + | ==Jump to Gene / Search FlyBase== |
| − | Jump to Gene (J2G) | + | Jump to Gene (J2G) and Search Flybase are alternative query tools found in the top-right of the blue navigation bar on every page in FlyBase - these allow for targeted and wide searches of FlyBase, respectively. |
| − | You can | + | ===Jump to Gene=== |
| + | The Jump to Gene (J2G) mode is a NAVIGATION tool, not a search tool, and thus should be used when you know the FlyBase symbol, name or ID for your gene, and you simply want to go directly to its corresponding gene report. You can enter a gene symbol, gene fullname, annotation (CG/CR) ID or FBgn ID into the J2G box (e.g. amn, amnesiac, CG11937 or FBgn0086782). You can also enter gene symbol synonyms or add wildcards (*), though doing so increases the likelihood that non-unique results will be returned - if there is one and only one hit, J2G will take you to a report page; if there are multiple hits, J2G generates a hit list. Note that J2G does NOT search synonyms of fullnames. | ||
| − | J2G | + | J2G processes your query in the following order:<br /> |
| − | # | + | # Primary FlyBase ID (FBgn). Any hits? Return hit(s), end<br /> |
| − | # | + | # Symbol (case-sensitive). Any hits? Return hit(s), end<br /> |
| − | + | # Symbol synonym (case-sensitive). Any hits? Return hit(s), end<br /> | |
| − | # synonym (case-sensitive) | + | # Symbol synonym (case-insensitive). Any hits? Return hit(s), end<br /> |
| − | # synonym (case-insensitive) | + | # Full name (case-insensitive). Any hits? Return hit(s), end<br /> |
| − | # name (case- | + | # Secondary FlyBase ID. Any hits? Return hit(s), end<br /> |
| − | # | ||
If nothing found, return error page | If nothing found, return error page | ||
| − | + | This tool also works with FBids for non-gene entities (e.g. FBal0090485 or FBab0002363) and with allele symbols (e.g. amn[X8]). | |
| + | J2G navigates to D. melanogaster gene symbols by default - if you would like to navigate to a non-melanogaster gene, you need to use the unique, 4-letter species abbreviation, followed by a backslash, and then the gene symbol (e.g. Dpse\dpp), or use the respective FBgn identifier. | ||
| − | + | ===Search FlyBase=== | |
| + | The Search FlyBase mode is the same as the [http://{{flybaseorg}}/ QuickSearch - Search FlyBase] tab found at the HomePage. It performs a comprehensive search of text-searchable FlyBase data across all classes of reports and the results are displayed in the form a Hit List summarizing the matching records by data type. For example, a search for 'amn' retrieves the matching reports for Aberration, Allele, Anatomy Ontology, Clone, Dataset, Gene, etc... | ||
| − | + | By clicking on one or more of the data types in the hit list it will only display individual matches within those data types. Click on any of the individual hits to view the corresponding report page. | |
| + | |||
| + | 'Search FlyBase' entries allow wildcards (*) to broaden the query. 'Search FlyBase' entries also allow multiple terms: a Boolean 'AND' is used as default (e.g. ‘cnn cbs’ equals to 'cnn AND cbs'). Adding 'OR' between terms will find records that have one or another of a list of terms (e.g. ‘cnn OR cbs’). To exclude certain terms from the results, use the ‘-’ character as a prefix (e.g. ‘Parkinson -CG5680’). Finally, results can be specified to contain an exact phrase by surrounding the search term with double quotes (e.g. “SH3 domain”). | ||
==QuickSearch== | ==QuickSearch== | ||
| − | [http://{{ | + | The [http://{{flybaseorg}} QuickSearch] tool on the FlyBase home page allows searching across all FlyBase reports. Forms for searching across all data types or for searching specific types of data have been separated into ‘tabs’, arrayed at the top of the QuickSearch window. Use the "Simple" tab to search all FlyBase reports. Results are in the form of a hit list summarizing the matching records by data type. More limited searches are available in the remaining tabs. You can search for particular curated data classes, e.g. genes, alleles, aberrations, etc. (Data Class tab), Human Disease models (Human Disease tab), Orthologs (Orthologs tab), GAL4 and other drivers and reporters (GAL4 etc tab), Protein Domains (Protein Domains tab), Gene Expression (Expression tab), Gene Groups (Gene Groups tab), Phenotype (Phenotype tab), Gene Ontolgy (GO tab), and References (References tab). |
| − | QuickSearch | + | QuickSearch searches ''D. melanogaster'' data by default. The "Simple, Expression, and "Data Class" tabs offer the option to search all species. If you want to search for a gene in a particular species, you can use the unique, 4-letter [[FlyBase:Abbreviations | species abbreviation]], followed by a backslash, and then the gene symbol (e.g. Dpse\dpp). |
| − | + | For a full description of the QuickSearch tabs, see [https://wiki.flybase.org/wiki/FlyBase:QuickSearch QuickSearch Help Page]. | |
| − | |||
| − | |||
| − | |||
| − | QuickSearch | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
==QueryBuilder== | ==QueryBuilder== | ||
| − | [http://{{ | + | [http://{{flybaseorg}}/cgi-bin/qb.pl QueryBuilder] (QB) provides the most powerful way to search FlyBase on a field-by-field level. QB presents a simple user interface that supports powerful searches by offering access to DataSet|Field pairs (for example, Genes|CV:GO:Molecular Function) in FlyBase along with the ability to include any combination of datasets in the same search (Note that Human Disease, Cell Line, Gene Group, and Strain reports are not yet accessible with QueryBuilder). QB automatically creates sets of records that are cross-referenced to the records that match your query, providing links to all related records in FlyBase from a single page. Both simple and complex queries can be built in a few steps. A search can be focused on a particular piece of data within a report page, such as the 'mapped features and mutations' associated with a gene, and Boolean operators (and, or, but not) can be used to combine two or more searches. QB allows a user to perform much more sophisticated searches compared to [http://{{flybaseorg}}/ QuickSearch] or other search tools on FlyBase, by taking full advantage of how the data is stored in FlyBase. A useful feature of QueryBuilder is that sets of results can be exported to QB from hitlists, as described in the 'Hit list refinement' section, and then modified to refine the search by adding additional query segments. Thus, QB is a very powerful tool that can be used in many different ways to explore the data in FlyBase. |
| − | The 'Query Builder Help' section on the '[http://{{ | + | The 'Query Builder Help' section on the '[http://{{flybaseorg}}/cgi-bin/qb.pl QueryBuilder Home Page]' outlines the basic search strategy. There are three options on the QB start page: select a pre-constructed query, import a previously saved query, and build a new query. Help for all of these options is available further down the page as well as a description of how to carry out an expression data search. |
==Vocabularies (previously known as TermLink)== | ==Vocabularies (previously known as TermLink)== | ||
| − | The [http://{{ | + | The [http://{{flybaseorg}}/vocabularies Vocabularies Search Page] provides easy access to data annotated with a particular controlled term or one of its synonyms. For example, you can use Vocabularies to retrieve a list of all the genes annotated with a particular GO term, or all the transcripts expressed in a particular body part. You do not need to know the precise term that FlyBase uses to store the data; the search box on the Vocabularies page retrieves controlled vocabulary terms that contain your query or terms that list a synonym containing the search term. For example, if you enter wing you will obtain a list that includes the controlled terms wing, anterior wing margin, and dorsal mesothoracic disc, which has the synonym wing disc. The controlled terms in the list are hyperlinked to TermReport pages that describe a single term in detail. Alternatively, you can also browse various controlled vocabulary hierarchies, by using the trees displayed on the main Vocabularies page. |
| − | The [http://{{ | + | The [http://{{flybaseorg}}/vocabularies Vocabularies Search] is the only search tool in FlyBase that allows users to search directly for controlled vocabulary (CV) term reports from any of the controlled vocabularies (CVs) used by FlyBase. This includes the GO and anatomy hierarchies, among others. Wildcards are automatically added to the beginning and the end of a search term. For each search performed, Vocabularies returns a hit list of CV term reports that match the search term. These are listed according to CV type, in the following order: anatomy term reports, FlyBase controlled vocabulary term reports, development term reports, GO term reports and SO term reports. Each term report allows the user to retrieve gene, allele, transcript, polypeptide or image reports associated with the term. |
| − | Please go to [ | + | Please go to [[FlyBase:Vocabularies | Vocabularies Help]] for more information. This page can also be accessed from the bottom of the [http://{{flybaseorg}}/vocabularies Vocabularies Search Page]. |
| − | There is a [https://www.youtube.com/watch?v= | + | There is a [https://www.youtube.com/watch?v=jgCON15SiRo video tutorial] on YouTube. |
=Query Results Analysis Tools= | =Query Results Analysis Tools= | ||
==HitList Refinement== | ==HitList Refinement== | ||
| − | When you perform any search that returns multiple hits, you are presented with a hit list, that can be modified or refined. By default all records are selected for inclusion in subsequent manipulations, but the checkboxes allow user-defined subsets to be created. | + | When you perform any search that returns multiple hits, you are presented with a hit list, that can be modified or refined. By default all records are selected for inclusion in subsequent manipulations, but the checkboxes allow user-defined subsets to be created. In Table mode, the first data column links directly to the report for each record that matched your search. Other columns link to [http://flybase.org/jbrowse/ JBrowse] or to searches that return hits directly related to that record. In addition to these links, the hit list provides a set of powerful tools for query refinement or batch processing. |
| − | The ' | + | The 'Convert' drop down menu enables you to see all objects of a particular class that are related to the hits selected in your list. For example, selecting 'clones' from the 'Convert' menu of a gene search will return a list of clones that are related to the selected genes. |
| − | The ' | + | The 'Analyze' button allows you to see the frequency of values within your selected hits for a predefined list of fields. Selecting 'Biological process', for example, from the Analyze tool for a list of genes involved in the Notch signalling pathway will result in a page listing the distribution of the different biological process controlled vocabulary terms associated with the list. Clicking on the number in the 'Related records' column in this new table will return the genes from your hitlist that are annotated to be involved in that GO term. |
| − | Lastly, the ' | + | Lastly, the 'Export' button allows you to send the selected hits to our Batch Download tool for use offline, to a new QueryBuilder session for further querying, or to link-out HTML tables of various third party data sources with data linked to the hits in your result list. The 'Ribbon Stack Viewer' allows the comparision of up to 100 genes using the Gene Ontology (GO) summary ribbon (as described [[FlyBase:Gene_Group_Report#GO_ribbon_stack|here]]). |
==Batch Download== | ==Batch Download== | ||
| − | The [http://{{ | + | The [http://{{flybaseorg}}/batchdownload Batch Download] tool provides bulk access to a variety of data for a specified list of unique IDs (please note: secondary IDs, synonyms, or full names are not allowed because they are not unique). |
IDs can be sent from a FlyBase hit list, uploaded from a local file, or entered manually. | IDs can be sent from a FlyBase hit list, uploaded from a local file, or entered manually. | ||
| − | The | + | The tool provides access to two types of data: data from specific fields in our web reports and data from our diverse collection of precomputed flat files. Any line from a precomputed file that matches the lists of IDs supplied can be downloaded using the precomputed file option. |
| + | |||
| + | The HTML table option allows you to create a custom report with only the fields you want while preserving hyperlinks for direct navigation to other FlyBase data. | ||
| + | |||
| + | ==ID Validator== | ||
| + | |||
| + | This tool will accept a list of FlyBase symbols/IDs (for any data type) and, where necessary/possible, update them to their current versions. It will also convert certain external IDs (GenBank nucleotide/protein accessions, UniProt accessions, PubMed IDs) into their equivalent FlyBase IDs. The output is provided as a validation table that can either be downloaded as a file or exported to a FlyBase HitList for futher processing (including conversion between data types). | ||
| − | + | For full details, see the [[FlyBase:ID_Validator|ID_Validator]] page. | |
=Genomic Search Tools and Browsers= | =Genomic Search Tools and Browsers= | ||
==BLAST== | ==BLAST== | ||
| − | [http://{{ | + | [http://{{flybaseorg}}/blast/ BLAST] (Basic Local Alignment Search Tool), provides a method for rapid searching of nucleotide and protein databases. FlyBase BLAST allows the opportunity to BLAST query the 12 completed Drosophila genomes, along with related insect species for which full genomes have been sequenced. BLAST provides access to the FASTA sequences of all sequenced Drosophila sequences, as well as providing links to GenBank. In addition, you can BLAST an unknown sequence and identify its position on JBrowse. |
| − | The [http://{{ | + | The [http://{{flybaseorg}}/blast/ BLAST] homepage is split into three sections; the first allows the user to input the query sequence and set-up the standard BLAST parameters (e.g. Expectation value, database to be searched); the second section allows the species to be selected; while the third allows the user to specify advanced BLAST options. |
Clicking on the hyperlinks provides hints and tips for the BLAST search. | Clicking on the hyperlinks provides hints and tips for the BLAST search. | ||
| − | == | + | ==JBrowse== |
| − | FlyBase [http://{{ | + | FlyBase [http://{{flybaseorg}}/jbrowse/?data=data%2Fjson%2Fdmel&loc=2R%3A13437075..13452700&tracks=Gene%2Cprotein_domains%2CcDNA&highlight= JBrowse] provides a graphical representation of the Drosophila melanogaster genome. JBrowse was developed by the Generic Model Organism Database ([http://gmod.org/wiki/Main_Page GMOD]) consortium and is the successor to GBrowse. |
| − | |||
| − | + | Genes, cDNAs, insertions, deficiencies, mapped mutations, regulatory features, RNAi reagents, RNA-seq data, and a wide array of other mapped features can be selected and viewed along a genome coordinate scale. You can navigate to a specific location by entering a precise sequence range, any valid FlyBase identifier for a gene, gene product, or insertion, or a cytological band in the 'Landmark or Region' box. Then move laterally along the genome by using the arrows at the top of the browser or by clicking in an open area of the viewer and dragging side to side. You can zoom in and out by clicking the plus and minus icons in the navigation bar or zoom in by selecting a region of the lower coordinate scale. You can move to a different region of the chromosome arm by clicking on a spot on the chromosome scale at the top of the viewer and switch to a different chromosome by using the chromosome selector at the top. | |
| − | + | FlyBase presents a view of D.melanogaster that displays gene models and the modENCODE Developmental stage RNA-seq track. Additional tracks can be selected from the 'Available Tracks' menu at the left side of the Browser. Tracks can be easily reordered by clicking on the track name and dragging to a new location on the viewer. Descriptions of individual tracks can be found in the [[FlyBase:JBrowse Tracks|FlyBase JBrowse Tracks]] document at the FlyBase wiki. | |
| − | + | See the [[FlyBase:JBrowse|FlyBase JBrowse Help]] wiki page for FlyBase-specific tips. More generic JBrowse help can be accessed from the Help menu in the upper row of the JBrowse page. | |
| − | + | ==Chromosome Maps== | |
| + | The [http://{{flybaseorg}}/maps/chromosomes/maps chromosome maps] show sequence scaffolds aligned to polytene chromosome maps for the Muller elements of the sequenced Drosophila species. For more information on the syntenic relationships among the 12 sequenced genomes, their standard chromosomal numbering and corresponding Muller element please see the [http://{{flybaseorg}}/maps/synteny Muller Element Arm Synteny] Table. | ||
| − | == | + | ==CytoSearch== |
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | A [http://{{flybaseorg}}/cytosearch CytoSearch] query will return lists of all the mapped genes, transgene insertions, and aberrations that are within, overlap or include the query region. The query returns features mapped both at the cytological and at the sequence level. Each hit includes the cytology, the observed (in green) or estimated (in red) sequence coordinates, and the symbol of the mapped feature as well as available stocks. | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | For CytoSearch searches, sequence-based data trumps cytology when both are available, cytology trumps meiotic data when both are available, and estimated cytology is used when only meiotic data are available. The FlyBase [[Media:DmelMapTable.160615c.xlsx|correspondence table spreadsheet]] for cytological and sequence level maps are used to estimate cytology from sequence range and sequence range from cytology, for both the underlying data and the query input. | |
| − | |||
CytoSearch is useful for searching for genetic objects mapped to a particular genomic region (but not necessarily mapped to the sequence). | CytoSearch is useful for searching for genetic objects mapped to a particular genomic region (but not necessarily mapped to the sequence). | ||
==Coordinate Converter== | ==Coordinate Converter== | ||
| − | The [http://{{ | + | The [http://{{flybaseorg}}/convert/coordinates Coordinate Converter] allows you to convert genomic coordinates between different genome releases. Just select the species, the input and output assemblies, enter your list of coordinates (or load them from a file), and away you go! It's that simple. The output page or file will list the input coordinates (with the release), the output coordinates (with the release), and notes on the conversion (e.g., "includes 1 area of change; different scaffold"). See the [[FlyBase:Coordinate_Converter | Coordinate Converter]] help page for more information. |
| − | == | + | ==Feature Mapper== |
| − | [http://{{ | + | The [http://{{flybaseorg}}/featuremapper Feature Mapper] tool allows you to do a search with one or many genes, sequence-based features or genomic regions and returns a wide variety of sequence-based genomic features that overlap or map within the associated genomic region(s). The reported features include gene structure features, aligned evidence, noncoding features, mapped mutations, and RNAi reagents. The search returns lists of features that map to the region(s) of interest. Enter the symbols or IDs for genomic features or a sequence region, check the features types that you wish to have returned, and submit your query. Search results can be saved as a GFF file or exported to a hitlist. For more information, see [[FlyBase:Feature Mapper | Feature Mapper Help]]. |
| + | |||
| + | ==Sequence Downloader== | ||
| + | The [http://{{flybaseorg}}/download/sequence/ Sequence Downloader] tool provides access to sequence data in FASTA format by ID or genomic location. This tool offers 3 modes of operation ID, Bulk ID, and Bulk Region. These modes can be toggled by using the '''Mode''' drop down option at the top of the tool. ID and Bulk ID modes accept IDs for genes (FBgn), transcripts (FBtr), polypeptides (FBpp), clones (FBcl), sequence features (FBsf), and recombinant constructs (FBtp). | ||
| + | |||
| + | [[File:Sequence_Downloader.png|thumb|Sequence Downloader]] | ||
| + | |||
| + | Some features of Sequence Downloader include: | ||
| − | + | * Viewing/downloading sequence by its ID (e.g. a gene by its FBgn ID). | |
| + | * Viewing/downloading sequence by its ID and subtype (e.g. all 5' UTRs of a gene). | ||
| + | * Showing the relative coordinates of a selected subregion of the sequence. | ||
| + | * Searching the sequence for a specific pattern with regular expressions. | ||
| + | * Viewing/downloading IDs in bulk. | ||
| + | * Viewing/downloading IDs by genomic location. | ||
| + | * Ability to add upstream/downstream bases to the specified genomic locations. | ||
| + | * Download sequence data of either strand. | ||
| − | + | See the full [[FlyBase:SequenceDownloader|Sequence Downloader docs]] for more information on how to use this tool. | |
| − | |||
| − | |||
| − | + | =RNA-Seq Query Tools and Browsers= | |
| − | + | The primary RNA-Seq data in FlyBase are the modENCODE data originally published in [http://{{flybaseorg}}/reports/FBrf0213330.html Graveley et al., 2011] and [http://{{flybaseorg}}/reports/FBrf0225793.html Brown et al., 2014], comprising 30 developmental stage expression profiles, 29 tissue expression profiles, 25 treatment/condition expression profiles and 24 cell line expression profiles. RNA-Seq reads were mapped to the Release 6 genome assembly as described in [http://{{flybaseorg}}/reports/FBrf0226107.html Brown et al., 2014]. In JBrowse genomic views, several other RNA-Seq datasets are also presented. The RNA-Seq query tools are restricted to the modENCODE datasets. For each modENCODE RNA-Seq sample, gene expression level was calculated as RPKM within the exonic extent of the gene, as described in [http://{{flybaseorg}}/reports/FBrf0221009.html Gelbart and Emmert, 2013]. For purposes of presentation and queries, values were assigned to one of eight bins, from very low to extremely high. | |
| − | [http://{{ | ||
| − | + | The available RNA-Seq query tools and browsers are listed below and described in more detail at the [[FlyBase:RNA-Seq Overview | RNA-Seq overview page]]. The direct link at the top of the FlyBase home page (top array of icons) also goes to this overview page. | |
| − | < | + | A series of '''video tutorials''' describing different RNA-Seq tools is available. See |
| + | *[https://www.youtube.com/watch?v=KpJVkopUBDM&t=107s RNA-Seq Part I: Using GBrowse] -- <span style="color:red"> Warning: The GBrowse genome viewer has been updated to JBrowse, but techniques shown here are still useful in JBrowse.</span> | ||
| + | *[https://www.youtube.com/watch?v=Ho_PZ4XB8y8&t=17s RNA-Seq Part II: Using RNA-Seq Profile Search] | ||
| + | *[https://www.youtube.com/watch?v=mcMpRxeX-KY RNA Seq Part III: Searching for Similarly Expressed Genes] | ||
| − | + | ==JBrowse== | |
| − | + | Use [http://{{flybaseorg}}/jbrowse/?data=data/json/dmel JBrowse] to view multiple RNA-Seq expression profiles across the genome. | |
| − | [http://{{ | ||
| − | + | ==RNA-Seq Profile== | |
| − | + | [http://{{flybaseorg}}/rnaseq/profile_search RNA-Seq Profile] is a fine grained query tool, powered by modENCODE high-throughput RNA-Seq expression data, that allows you to find genes with specific patterns of expression across several variables. | |
| − | + | Go to [https://www.youtube.com/watch?v=Ho_PZ4XB8y8&t=17s RNA-Seq Part II: Using RNA-Seq Profile Search] to see the associated video tutorial or to [[FlyBase:RNA-Seq Overview | RNA-Seq Overview]] for more information on using the tool. | |
| − | = | + | ==RNA-Seq Similarity== |
| + | |||
| + | Use [http://{{flybaseorg}}/rnaseq/simsearch RNA-Seq Similarity] to find genes with expression patterns that are similar to that of a given gene; this search option can also be launched from the relevant gene page. | ||
| − | + | Go to [https://www.youtube.com/watch?v=mcMpRxeX-KY RNA Seq Part III: Searching for Similarly Expressed Genes] to see the associated video tutorial or to [[FlyBase:RNA-Seq Overview | RNA-Seq Overview]] for more information on using the tool. | |
| − | |||
| − | == | + | ==RNA-Seq By Region== |
| − | |||
| − | + | Use [http://{{flybaseorg}}/rnaseq/region RNA-Seq By Region] to compare the RNA-Seq signal for a given region across samples, or to compare signal between two regions within a single sample. | |
| − | [http://{{ | ||
| − | + | See [[FlyBase:RNA-Seq Overview | RNA-Seq Overview]] for more information on using the tool. | |
| − | FlyBase | ||
| − | + | =Other Tools= | |
| − | + | ==Interactions Browser== | |
| + | The [http://{{flybaseorg}}/cgi-bin/get_interactions.pl Interactions Browser] is accessible under the 'Tools' menu, at the top of the Interaction section of allele reports, or at the top of physical interaction reports. This tool provides a graphical way of exploring physical interaction data, or genetic interaction data (enhancer data only, suppressor data only, or both). For genetic interactions, the browser works in two modes: you can either search for the interactions of an allele, or the interactions of a gene. The latter will show the interactions of all alleles of the gene. Each node of an interaction diagram is a hyperlink, which enables you to navigate and browse the complex web of known genetic or physical interactions. Placing your cursor over the center of a node activates a pop-up window that in the case of a network of gene interactions contains a summary of the function of that particular gene, while in the case of interactions between alleles shows the context in which the interactions of that allele have been reported. For more information, go to the [http://wiki.flybase.org/wiki/FlyBase:Interaction_Browser Interactions Browser help documentation]. | ||
| − | + | ==ImageBrowse== | |
| + | [http://flybase.org/imagebrowse/ ImageBrowse] allows the user to browse through [https://wiki.flybase.org/wiki/FlyBase:Image_Report image reports] by organ system, life-cycle, tagma, or germ layer, as well as to browse images of different Drosophilids. This section also gives access to [http://flybase.org/imagebrowse/posters posters of common visible markers in D. melanogaster], as well as miscellaneous images and quick-time films. [http://{{flybaseorg}}/wiki/FlyBase:Controlled_vocabularies_used_by_FlyBase Controlled vocabulary terms] are used to annotate and label the images. To search images, and to link relevant gene, allele, transcript and protein records to stages of development, a region of the body or to a specific body part, go to [http://{{flybaseorg}}/vocabularies Vocabularies]. | ||
| − | == | + | ==Fast-Track Your Paper== |
| + | The [http://{{flybaseorg}}/submission/publication/ Fast-Track Your Paper] tool allows first-pass curation by users, indicating to curators the types of data in the paper and also resulting in relevant genes being associated with the reference. Corresponding authors of new publications are emailed and asked to use FTYP when their paper has been fully published (i.e. has final volume and page numbers) and has been added to our database. | ||
| − | + | Please go to the [[FlyBase:Fast Track Your Paper | Fast Track Your Paper]] page for more information. | |
| − | |||
[[Category:FlyBase]] [[Category:Help]] | [[Category:FlyBase]] [[Category:Help]] | ||
Latest revision as of 08:57, 7 June 2024
General Search Help and Tips
FlyBase can be searched for genes, alleles, aberrations and other genetic objects, phenotypes, sequences, stocks, images and movies, controlled terms, and Drosophila researchers using the tools available from the 'Tools' drop-down menu in the Navigation bar. In addition to the Navigation bar, which can be accessed from any FlyBase page, the homepage also has direct links to the most commonly used tools.
Below are brief descriptions of each of the tools, which have been split into five main sections:
- Overview of Search Strategies (for example, how to search for expression data)
- Main Query Tools (Jump to Gene, QueryBuilder, etc.)
- Query Results Analysis Tools (Hit list refinement, Batch Download)
- Genomic Search Tools and Browsers (JBrowse, BLAST etc.)
- Other Tools (Interactions Browser, Fast Track Your Paper etc.)
Links to Full documentation for FlyBase tools can be found in the Tools section of the FlyBase Help Index.
Overview of Search Strategies
Searching 12 species
Please note -- Starting in 2018, FlyBase will reflect updated gene models annotated by the NCBI gnomon pipeline for four species only: D. simulans, D. ananassae, D. pseudoobscura, and D. virilis. Thus, existing gene records for the other seven AAA species may go stale and newly annotated genes will not be included. (D. melanogaster gene models are updated via a separate pipeline.) This section will be updated as these changes occur.
Individual gene reports for genes from the 12 originally sequenced Drosophila genomes are available in FlyBase. There are four main ways in which these data can be browsed and queried in FlyBase:
- Gene Report Pages
For those interested in genome-wide analyses, bioinformatics and comparative genomics, there are a selection of pre-computed files available for download from our precomputed files page (in the Genomes:Annotation and Sequence section, for example), found in the 'Files' menu.
For those with an interest in a specific gene/protein/region across the different species, there are a number of ways to query the data. Our BLAST server allows querying of numerous sequenced insect genomes, either individually, as a subset, or all together.
Aberrations - deficiencies, dupications, inversions, translocations
One of the problems in a field of the size and complexity of Drosophila genetics is the use of nomenclature. This can lead to a number of names being given to the same object, and to the valid FlyBase name or symbol of an object being quite confusing or indeed not in common lab parlance. Aberration naming is no exception. The simplest ways to search for an aberration are either using CytoSearch, when you want to find an aberration that removes a particular gene or uncovers a cytological band, or using QuickSearch (use the 'Data Class' tab and select 'aberration' as the data class). Remember to use wildcards (i.e. *) to allow for slight differences in naming. FlyBase records all mentions of an aberration, so if an aberration is given a particular symbol in a paper, this name will be recorded as a synonym of the FlyBase 'valid' symbol (see the nomenclature document for more details). Alternatively, you can browse the molecularly localized aberrations for each chromosome by scanning JBrowse after selecting all "Aberrations" tracks.
Cytologically Mapped Features
When looking for cytology, you have a choice of a number of tools on FlyBase, including QueryBuilder. The easiest tools to use however, are CytoSearch or JBrowse. JBrowse is especially useful when looking for molecularly mapped sequences, insertions, or probes. CytoSearch comes into its own when searching for cytologically defined features, such as cytologically-mapped genes or deficiencies, that haven't been molecularly mapped to the sequence. Of course, as with many aspects of research, complimentary methods should be used. Therefore, we recommend you use both JBrowse and CytoSearch to analyse cytology.
A description of how FlyBase computes the cytological location for features that have been mapped to the genome can be found in Computed cytological data. Illustrations or electron micrographs of D. melanogaster polytene chromosomes as well as a cytogenetic-genetic-sequence location correspondence table can be found at D. melanogaster Chromosome Maps.
Expression Data
Browsing Expression Data
Expression patterns are captured by FlyBase curators for transcripts, proteins, and "reporters" (i.e. enhancer trap insertions and reporter constructs). Information about transcript and protein expression patterns can be found on gene reports (e.g. the elav gene), data for reporter constructs can be found on recombinant construct reports (e.g. P{elav-lacZ.H}) and associated allele reports (e.g. Ecol\lacZelav.PH), and data for enhancer or protein traps can be found on insertion reports (e.g. P{GawB}elavC155) and associated allele reports (e.g. Scer\GAL4elav-C155). In all cases, expression data will be found in the "Expression Data" section of the report. For those constructs or insertions that reflect expression of a particular gene, data are also promoted to the corresponding gene report, in a subsection of "Expression Data" labeled "Expression Deduced from Reporters" (e.g. expression data for both P{elav-lacZ.H} and P{GawB}elavC155 are displayed on the elav gene report). Subcellular Localization of protein is populated from Gene Ontology (GO) Cellular Component curation of genes.
We cooperate with several other databases of expression data and either display a portion of their data within FlyBase (e.g. FlyExpress) and/or link to their database (e.g. Fly-FISH). These types of data can be found in the "External Data & Images" subsection of the "Expression Data" section. Additionally, we maintain a set of links to Image Based Resources, including image databases, tools for image analysis, and tools for image visualization and annotation.
High throughput expression data from FlyAtlas and modENCODE can be found on Gene Reports in a subsection of "Expression Data" labeled "High-Throughput Expression Data". The modENCODE data can be visualized as a linear or log graph, or as a heatmap. The FlyAtlas section also includes a 'back-to-back' option, in which gene expression levels in larval tissues are juxtaposed with gene expression levels in the corresponding adult tissues. The graph displays can be scaled by gene maximum expressed, or by low, moderate, or high expression bin max.
Searching for Expression Patterns
Expression data curated from literature can be searched most easily and accurately by using the QuickSearch Expression or Gal4 etc tabs (detailed help at links). The Expression tab allows searches for genes by temporal-spatial expression pattern, while the GAL4 etc allows searches for GAL4 and other binary drivers, and non-binary reporters. Another expression pattern search option is QueryBuilder, which supports multipart queries (e.g. generate a list of genes which have the GO term "transcription factor activity" and whose protein products are expressed in the central nervous system). However, if you're interested in all genes expressed in a bodypart, tissue, or developmental stage, you can find that using Vocabularies. For example, by entering the term "adult mushroom body" into Vocabularies, you can obtain a list of genes expressed in that tissue.
Searching for High-Throughput Expression Patterns
RNA-Seq expression data can be searched to identify genes with specific expression characteristics using the RNA-Seq Profile Search tool. Genes that have expression patterns similar to a given gene can be found using the RNA-Seq Similarity Search tool; this search option can also be launched from the relevant gene page. RNA-Seq By Region can be used to compare the RNA-Seq signal for a given region across samples, or to compare signal between two regions within a single sample. For additional information about these tools see RNA-Seq Query Tools and Browsers.
Mutant Phenotype Data
Mutant phenotype data is associated with alleles in FlyBase, so you need to search allele data if you are interested in mutant phenotype. In addition to free text describing the phenotype, the alleles are indexed with controlled vocabulary (CV) terms, which makes it easier for you to search for a particular phenotype, e.g. searching for mutants that affect the wing. You can search with these CV terms using either Vocabularies or QueryBuilder.
You can find mutant alleles affecting the wing from all species using Vocabularies. If you enter the term "wing" into Vocabularies search page and then click on the "Alleles" button in the report page, you will obtain a list of mutant alleles that affect the wing. However, to search in a specific species, or to search for mutant phenotypes as part of a multipart query, QueryBuilder must be used. In this case, you should pick the "CV Hierarchy (GO/etc.)" dataset and then use the term picker to choose the body part, e.g. wing. In both cases, the default is to search both for alleles specifically labelled with the CV term, e.g. wing and also with child CV terms that are a subset of the term chosen, e.g. wing vein. If you want to restrict your search to just the precise term chosen, use QueryBuilder and select 'Retrieve records annotated with "This CV term only"' before you run the query.
Find a related video tutorial at Finding genes with similar phenotypes.
References
FlyBase is an excellent source of Drosophila references. References can be searched in a number of ways. The easiest way is through QuickSearch, on our homepage. Choose the 'References' tab and fill in one or more of the search boxes. The field identity of each search box can be modified using the dropdown menus at the left. For more information, please go to the QuickSearch Help Page.
More refined reference searches can be performed using QueryBuilder (QB). Click on the box titled 'Query is empty.. Click here to start building' on the QB start page to being the search. At this stage the window will be displaying all the fields available to search for the 'Genes' dataset. Change the dataset to 'References'. Now the fields found in the reference reports are displayed. From here, you can search all the data found in the reference report, including pubmed ID, author, and type (e.g. review). Find complete instructions on the QueryBuilder tool page.
A popular way to search for references is to search for a (list of) objects (e.g. genes, GO terms) and then to use the 'Show related' toggle on the hits page to change the hit list to the related references. The 'Results Analysis/Refinement' button, found on the hit list page, can be used to analyse the distribution of the references over year, journal, author, and type of publication (e.g. review, paper, abstract).
Stocks
One of the easiest ways to search for a stock in FlyBase is to use QuickSearch. Simply change the data class to 'stocks', type in the feature of interest (e.g. a gene symbol, allele symbol), and search. A further way to identify stocks is through the hit list produced after a search. At the top of the hit list there is a toggle allowing you to 'Show related' stocks. Stocks can also be found for individual alleles by clicking on the Stocks matryoska on the allele report page.
Main Query Tools
Jump to Gene / Search FlyBase
Jump to Gene (J2G) and Search Flybase are alternative query tools found in the top-right of the blue navigation bar on every page in FlyBase - these allow for targeted and wide searches of FlyBase, respectively.
Jump to Gene
The Jump to Gene (J2G) mode is a NAVIGATION tool, not a search tool, and thus should be used when you know the FlyBase symbol, name or ID for your gene, and you simply want to go directly to its corresponding gene report. You can enter a gene symbol, gene fullname, annotation (CG/CR) ID or FBgn ID into the J2G box (e.g. amn, amnesiac, CG11937 or FBgn0086782). You can also enter gene symbol synonyms or add wildcards (*), though doing so increases the likelihood that non-unique results will be returned - if there is one and only one hit, J2G will take you to a report page; if there are multiple hits, J2G generates a hit list. Note that J2G does NOT search synonyms of fullnames.
J2G processes your query in the following order:
- Primary FlyBase ID (FBgn). Any hits? Return hit(s), end
- Symbol (case-sensitive). Any hits? Return hit(s), end
- Symbol synonym (case-sensitive). Any hits? Return hit(s), end
- Symbol synonym (case-insensitive). Any hits? Return hit(s), end
- Full name (case-insensitive). Any hits? Return hit(s), end
- Secondary FlyBase ID. Any hits? Return hit(s), end
If nothing found, return error page
This tool also works with FBids for non-gene entities (e.g. FBal0090485 or FBab0002363) and with allele symbols (e.g. amn[X8]). J2G navigates to D. melanogaster gene symbols by default - if you would like to navigate to a non-melanogaster gene, you need to use the unique, 4-letter species abbreviation, followed by a backslash, and then the gene symbol (e.g. Dpse\dpp), or use the respective FBgn identifier.
Search FlyBase
The Search FlyBase mode is the same as the QuickSearch - Search FlyBase tab found at the HomePage. It performs a comprehensive search of text-searchable FlyBase data across all classes of reports and the results are displayed in the form a Hit List summarizing the matching records by data type. For example, a search for 'amn' retrieves the matching reports for Aberration, Allele, Anatomy Ontology, Clone, Dataset, Gene, etc...
By clicking on one or more of the data types in the hit list it will only display individual matches within those data types. Click on any of the individual hits to view the corresponding report page.
'Search FlyBase' entries allow wildcards (*) to broaden the query. 'Search FlyBase' entries also allow multiple terms: a Boolean 'AND' is used as default (e.g. ‘cnn cbs’ equals to 'cnn AND cbs'). Adding 'OR' between terms will find records that have one or another of a list of terms (e.g. ‘cnn OR cbs’). To exclude certain terms from the results, use the ‘-’ character as a prefix (e.g. ‘Parkinson -CG5680’). Finally, results can be specified to contain an exact phrase by surrounding the search term with double quotes (e.g. “SH3 domain”).
QuickSearch
The QuickSearch tool on the FlyBase home page allows searching across all FlyBase reports. Forms for searching across all data types or for searching specific types of data have been separated into ‘tabs’, arrayed at the top of the QuickSearch window. Use the "Simple" tab to search all FlyBase reports. Results are in the form of a hit list summarizing the matching records by data type. More limited searches are available in the remaining tabs. You can search for particular curated data classes, e.g. genes, alleles, aberrations, etc. (Data Class tab), Human Disease models (Human Disease tab), Orthologs (Orthologs tab), GAL4 and other drivers and reporters (GAL4 etc tab), Protein Domains (Protein Domains tab), Gene Expression (Expression tab), Gene Groups (Gene Groups tab), Phenotype (Phenotype tab), Gene Ontolgy (GO tab), and References (References tab).
QuickSearch searches D. melanogaster data by default. The "Simple, Expression, and "Data Class" tabs offer the option to search all species. If you want to search for a gene in a particular species, you can use the unique, 4-letter species abbreviation, followed by a backslash, and then the gene symbol (e.g. Dpse\dpp).
For a full description of the QuickSearch tabs, see QuickSearch Help Page.
QueryBuilder
QueryBuilder (QB) provides the most powerful way to search FlyBase on a field-by-field level. QB presents a simple user interface that supports powerful searches by offering access to DataSet|Field pairs (for example, Genes|CV:GO:Molecular Function) in FlyBase along with the ability to include any combination of datasets in the same search (Note that Human Disease, Cell Line, Gene Group, and Strain reports are not yet accessible with QueryBuilder). QB automatically creates sets of records that are cross-referenced to the records that match your query, providing links to all related records in FlyBase from a single page. Both simple and complex queries can be built in a few steps. A search can be focused on a particular piece of data within a report page, such as the 'mapped features and mutations' associated with a gene, and Boolean operators (and, or, but not) can be used to combine two or more searches. QB allows a user to perform much more sophisticated searches compared to QuickSearch or other search tools on FlyBase, by taking full advantage of how the data is stored in FlyBase. A useful feature of QueryBuilder is that sets of results can be exported to QB from hitlists, as described in the 'Hit list refinement' section, and then modified to refine the search by adding additional query segments. Thus, QB is a very powerful tool that can be used in many different ways to explore the data in FlyBase.
The 'Query Builder Help' section on the 'QueryBuilder Home Page' outlines the basic search strategy. There are three options on the QB start page: select a pre-constructed query, import a previously saved query, and build a new query. Help for all of these options is available further down the page as well as a description of how to carry out an expression data search.
Vocabularies (previously known as TermLink)
The Vocabularies Search Page provides easy access to data annotated with a particular controlled term or one of its synonyms. For example, you can use Vocabularies to retrieve a list of all the genes annotated with a particular GO term, or all the transcripts expressed in a particular body part. You do not need to know the precise term that FlyBase uses to store the data; the search box on the Vocabularies page retrieves controlled vocabulary terms that contain your query or terms that list a synonym containing the search term. For example, if you enter wing you will obtain a list that includes the controlled terms wing, anterior wing margin, and dorsal mesothoracic disc, which has the synonym wing disc. The controlled terms in the list are hyperlinked to TermReport pages that describe a single term in detail. Alternatively, you can also browse various controlled vocabulary hierarchies, by using the trees displayed on the main Vocabularies page.
The Vocabularies Search is the only search tool in FlyBase that allows users to search directly for controlled vocabulary (CV) term reports from any of the controlled vocabularies (CVs) used by FlyBase. This includes the GO and anatomy hierarchies, among others. Wildcards are automatically added to the beginning and the end of a search term. For each search performed, Vocabularies returns a hit list of CV term reports that match the search term. These are listed according to CV type, in the following order: anatomy term reports, FlyBase controlled vocabulary term reports, development term reports, GO term reports and SO term reports. Each term report allows the user to retrieve gene, allele, transcript, polypeptide or image reports associated with the term.
Please go to Vocabularies Help for more information. This page can also be accessed from the bottom of the Vocabularies Search Page.
There is a video tutorial on YouTube.
Query Results Analysis Tools
HitList Refinement
When you perform any search that returns multiple hits, you are presented with a hit list, that can be modified or refined. By default all records are selected for inclusion in subsequent manipulations, but the checkboxes allow user-defined subsets to be created. In Table mode, the first data column links directly to the report for each record that matched your search. Other columns link to JBrowse or to searches that return hits directly related to that record. In addition to these links, the hit list provides a set of powerful tools for query refinement or batch processing.
The 'Convert' drop down menu enables you to see all objects of a particular class that are related to the hits selected in your list. For example, selecting 'clones' from the 'Convert' menu of a gene search will return a list of clones that are related to the selected genes.
The 'Analyze' button allows you to see the frequency of values within your selected hits for a predefined list of fields. Selecting 'Biological process', for example, from the Analyze tool for a list of genes involved in the Notch signalling pathway will result in a page listing the distribution of the different biological process controlled vocabulary terms associated with the list. Clicking on the number in the 'Related records' column in this new table will return the genes from your hitlist that are annotated to be involved in that GO term.
Lastly, the 'Export' button allows you to send the selected hits to our Batch Download tool for use offline, to a new QueryBuilder session for further querying, or to link-out HTML tables of various third party data sources with data linked to the hits in your result list. The 'Ribbon Stack Viewer' allows the comparision of up to 100 genes using the Gene Ontology (GO) summary ribbon (as described here).
Batch Download
The Batch Download tool provides bulk access to a variety of data for a specified list of unique IDs (please note: secondary IDs, synonyms, or full names are not allowed because they are not unique).
IDs can be sent from a FlyBase hit list, uploaded from a local file, or entered manually.
The tool provides access to two types of data: data from specific fields in our web reports and data from our diverse collection of precomputed flat files. Any line from a precomputed file that matches the lists of IDs supplied can be downloaded using the precomputed file option.
The HTML table option allows you to create a custom report with only the fields you want while preserving hyperlinks for direct navigation to other FlyBase data.
ID Validator
This tool will accept a list of FlyBase symbols/IDs (for any data type) and, where necessary/possible, update them to their current versions. It will also convert certain external IDs (GenBank nucleotide/protein accessions, UniProt accessions, PubMed IDs) into their equivalent FlyBase IDs. The output is provided as a validation table that can either be downloaded as a file or exported to a FlyBase HitList for futher processing (including conversion between data types).
For full details, see the ID_Validator page.
Genomic Search Tools and Browsers
BLAST
BLAST (Basic Local Alignment Search Tool), provides a method for rapid searching of nucleotide and protein databases. FlyBase BLAST allows the opportunity to BLAST query the 12 completed Drosophila genomes, along with related insect species for which full genomes have been sequenced. BLAST provides access to the FASTA sequences of all sequenced Drosophila sequences, as well as providing links to GenBank. In addition, you can BLAST an unknown sequence and identify its position on JBrowse.
The BLAST homepage is split into three sections; the first allows the user to input the query sequence and set-up the standard BLAST parameters (e.g. Expectation value, database to be searched); the second section allows the species to be selected; while the third allows the user to specify advanced BLAST options.
Clicking on the hyperlinks provides hints and tips for the BLAST search.
JBrowse
FlyBase JBrowse provides a graphical representation of the Drosophila melanogaster genome. JBrowse was developed by the Generic Model Organism Database (GMOD) consortium and is the successor to GBrowse.
Genes, cDNAs, insertions, deficiencies, mapped mutations, regulatory features, RNAi reagents, RNA-seq data, and a wide array of other mapped features can be selected and viewed along a genome coordinate scale. You can navigate to a specific location by entering a precise sequence range, any valid FlyBase identifier for a gene, gene product, or insertion, or a cytological band in the 'Landmark or Region' box. Then move laterally along the genome by using the arrows at the top of the browser or by clicking in an open area of the viewer and dragging side to side. You can zoom in and out by clicking the plus and minus icons in the navigation bar or zoom in by selecting a region of the lower coordinate scale. You can move to a different region of the chromosome arm by clicking on a spot on the chromosome scale at the top of the viewer and switch to a different chromosome by using the chromosome selector at the top.
FlyBase presents a view of D.melanogaster that displays gene models and the modENCODE Developmental stage RNA-seq track. Additional tracks can be selected from the 'Available Tracks' menu at the left side of the Browser. Tracks can be easily reordered by clicking on the track name and dragging to a new location on the viewer. Descriptions of individual tracks can be found in the FlyBase JBrowse Tracks document at the FlyBase wiki.
See the FlyBase JBrowse Help wiki page for FlyBase-specific tips. More generic JBrowse help can be accessed from the Help menu in the upper row of the JBrowse page.
Chromosome Maps
The chromosome maps show sequence scaffolds aligned to polytene chromosome maps for the Muller elements of the sequenced Drosophila species. For more information on the syntenic relationships among the 12 sequenced genomes, their standard chromosomal numbering and corresponding Muller element please see the Muller Element Arm Synteny Table.
CytoSearch
A CytoSearch query will return lists of all the mapped genes, transgene insertions, and aberrations that are within, overlap or include the query region. The query returns features mapped both at the cytological and at the sequence level. Each hit includes the cytology, the observed (in green) or estimated (in red) sequence coordinates, and the symbol of the mapped feature as well as available stocks.
For CytoSearch searches, sequence-based data trumps cytology when both are available, cytology trumps meiotic data when both are available, and estimated cytology is used when only meiotic data are available. The FlyBase correspondence table spreadsheet for cytological and sequence level maps are used to estimate cytology from sequence range and sequence range from cytology, for both the underlying data and the query input.
CytoSearch is useful for searching for genetic objects mapped to a particular genomic region (but not necessarily mapped to the sequence).
Coordinate Converter
The Coordinate Converter allows you to convert genomic coordinates between different genome releases. Just select the species, the input and output assemblies, enter your list of coordinates (or load them from a file), and away you go! It's that simple. The output page or file will list the input coordinates (with the release), the output coordinates (with the release), and notes on the conversion (e.g., "includes 1 area of change; different scaffold"). See the Coordinate Converter help page for more information.
Feature Mapper
The Feature Mapper tool allows you to do a search with one or many genes, sequence-based features or genomic regions and returns a wide variety of sequence-based genomic features that overlap or map within the associated genomic region(s). The reported features include gene structure features, aligned evidence, noncoding features, mapped mutations, and RNAi reagents. The search returns lists of features that map to the region(s) of interest. Enter the symbols or IDs for genomic features or a sequence region, check the features types that you wish to have returned, and submit your query. Search results can be saved as a GFF file or exported to a hitlist. For more information, see Feature Mapper Help.
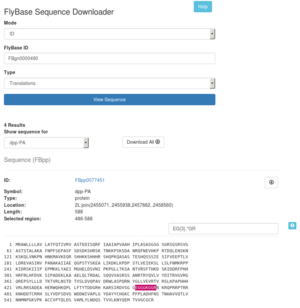
Sequence Downloader
The Sequence Downloader tool provides access to sequence data in FASTA format by ID or genomic location. This tool offers 3 modes of operation ID, Bulk ID, and Bulk Region. These modes can be toggled by using the Mode drop down option at the top of the tool. ID and Bulk ID modes accept IDs for genes (FBgn), transcripts (FBtr), polypeptides (FBpp), clones (FBcl), sequence features (FBsf), and recombinant constructs (FBtp).
Some features of Sequence Downloader include:
- Viewing/downloading sequence by its ID (e.g. a gene by its FBgn ID).
- Viewing/downloading sequence by its ID and subtype (e.g. all 5' UTRs of a gene).
- Showing the relative coordinates of a selected subregion of the sequence.
- Searching the sequence for a specific pattern with regular expressions.
- Viewing/downloading IDs in bulk.
- Viewing/downloading IDs by genomic location.
- Ability to add upstream/downstream bases to the specified genomic locations.
- Download sequence data of either strand.
See the full Sequence Downloader docs for more information on how to use this tool.
RNA-Seq Query Tools and Browsers
The primary RNA-Seq data in FlyBase are the modENCODE data originally published in Graveley et al., 2011 and Brown et al., 2014, comprising 30 developmental stage expression profiles, 29 tissue expression profiles, 25 treatment/condition expression profiles and 24 cell line expression profiles. RNA-Seq reads were mapped to the Release 6 genome assembly as described in Brown et al., 2014. In JBrowse genomic views, several other RNA-Seq datasets are also presented. The RNA-Seq query tools are restricted to the modENCODE datasets. For each modENCODE RNA-Seq sample, gene expression level was calculated as RPKM within the exonic extent of the gene, as described in Gelbart and Emmert, 2013. For purposes of presentation and queries, values were assigned to one of eight bins, from very low to extremely high.
The available RNA-Seq query tools and browsers are listed below and described in more detail at the RNA-Seq overview page. The direct link at the top of the FlyBase home page (top array of icons) also goes to this overview page.
A series of video tutorials describing different RNA-Seq tools is available. See
- RNA-Seq Part I: Using GBrowse -- Warning: The GBrowse genome viewer has been updated to JBrowse, but techniques shown here are still useful in JBrowse.
- RNA-Seq Part II: Using RNA-Seq Profile Search
- RNA Seq Part III: Searching for Similarly Expressed Genes
JBrowse
Use JBrowse to view multiple RNA-Seq expression profiles across the genome.
RNA-Seq Profile
RNA-Seq Profile is a fine grained query tool, powered by modENCODE high-throughput RNA-Seq expression data, that allows you to find genes with specific patterns of expression across several variables.
Go to RNA-Seq Part II: Using RNA-Seq Profile Search to see the associated video tutorial or to RNA-Seq Overview for more information on using the tool.
RNA-Seq Similarity
Use RNA-Seq Similarity to find genes with expression patterns that are similar to that of a given gene; this search option can also be launched from the relevant gene page.
Go to RNA Seq Part III: Searching for Similarly Expressed Genes to see the associated video tutorial or to RNA-Seq Overview for more information on using the tool.
RNA-Seq By Region
Use RNA-Seq By Region to compare the RNA-Seq signal for a given region across samples, or to compare signal between two regions within a single sample.
See RNA-Seq Overview for more information on using the tool.
Other Tools
Interactions Browser
The Interactions Browser is accessible under the 'Tools' menu, at the top of the Interaction section of allele reports, or at the top of physical interaction reports. This tool provides a graphical way of exploring physical interaction data, or genetic interaction data (enhancer data only, suppressor data only, or both). For genetic interactions, the browser works in two modes: you can either search for the interactions of an allele, or the interactions of a gene. The latter will show the interactions of all alleles of the gene. Each node of an interaction diagram is a hyperlink, which enables you to navigate and browse the complex web of known genetic or physical interactions. Placing your cursor over the center of a node activates a pop-up window that in the case of a network of gene interactions contains a summary of the function of that particular gene, while in the case of interactions between alleles shows the context in which the interactions of that allele have been reported. For more information, go to the Interactions Browser help documentation.
ImageBrowse
ImageBrowse allows the user to browse through image reports by organ system, life-cycle, tagma, or germ layer, as well as to browse images of different Drosophilids. This section also gives access to posters of common visible markers in D. melanogaster, as well as miscellaneous images and quick-time films. Controlled vocabulary terms are used to annotate and label the images. To search images, and to link relevant gene, allele, transcript and protein records to stages of development, a region of the body or to a specific body part, go to Vocabularies.
Fast-Track Your Paper
The Fast-Track Your Paper tool allows first-pass curation by users, indicating to curators the types of data in the paper and also resulting in relevant genes being associated with the reference. Corresponding authors of new publications are emailed and asked to use FTYP when their paper has been fully published (i.e. has final volume and page numbers) and has been added to our database.
Please go to the Fast Track Your Paper page for more information.