FlyBase:SequenceDownloader
Sequence Downloader
The Sequence Downloader tool provides access to sequence data by ID or genomic location. This tool offers 3 modes of operation ID, Bulk ID, and Bulk Region. These modes can be toggled by using the Mode drop down option at the top of the tool.
ID Mode
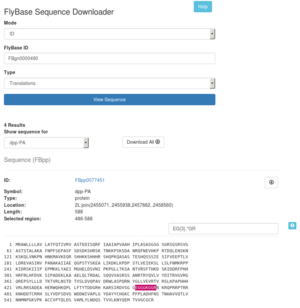
ID mode accepts IDs for genes (FBgn), transcripts (FBtr), polypeptides (FBpp), clones (FBcl), sequence features (FBsf), and recombinant constructs (FBtp). To view a sequence, enter in your ID, select the appropriate Type and click View Sequence. The sequence and some associated information will appear at the bottom of the page. This view offers several useful features for working with the sequence data. To download the sequence as a FASTA file locally, click the download icon in the top right of the sequence section.
Selecting a sequence region with your mouse, will display the range of your selection in the Selected region field.
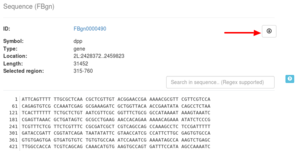
With the Search in sequence box, you can search your sequence for a specific pattern by entering the pattern in the input box. Matches will be highlighted within the sequence. This box also supports regular expression patterns. For more information see the small help icon next to the Search in sequence box.
This mode also allows you to easily download items that are associated with your desired ID. For example, when entering a FBgn ID, you can download all transcripts, polypeptides, UTRs, clones, etc. for that gene. When your ID and Type result in multiple sequences, the viewer will offer a drop down list from which you can choose the specific item to view. You can also download all items via the Download All button.
Bulk ID Mode
Bulk ID mode supports the display and downloading of sequence data for many IDs at once. Bulk ID mode supports IDs that are supported by ID mode. To view or download sequence data as a plain text FASTA file, enter in all your IDs, select your sequence type and click the Download button.
Bulk Region Mode
Bulk Region mode supports the downloading of sequence data by genomic location for any of the sequenced species in FlyBase. To download sequence by genomic location, select your species, enter in one or more coordinates (one per line), and click "Download". Click the small help icon next to Sequence Coordinates to see the allowed formats for the location strings. With the Additional flanking bases input box you can specify additional flanking bases (5' and 3') that will be added to all sequence locations; simply enter in the number of bases in the input box. Finally, the Strand option allows you to retrieve all sequences from either the plus or minus strand of all the sequence locations.
Links to Sequence Downloader
Sequence Downloader supports direct links (via GET or POST) from external sources into the tool itself.
Links to ID Mode
For ID mode you have to supply a valid FlyBase ID and a supported type. Supported types include FBgn, gene_extended, CDS, intron, exon, FBpp, FBtr, five_prime_utr, three_prime_utr, FBcl, FBsf, FBtp.
- http://flybase.org/download/sequence/[FlyBase ID]/[Type] (GET only)
- http://flybase.org/download/sequence/FBgn0000490
- http://flybase.org/download/sequence/FBgn0000490/CDS
- http://flybase.org/download/sequence/FBgn0000490/five_prime_utr
Links to Bulk ID Mode
For Bulk ID mode, you can pass in a comma delimited list of IDs, sequence type, and output format as parameters and it will populate the form with the specified values. See ID mode for a list of valid types.
- http://flybase.org/download/sequence/batch/?ids=[FlyBase IDs]&type=[Type]&output=[Output] (GET OR POST)
- http://flybase.org/download/sequence/batch/?ids=FBgn0000490,FBgn0013765
- http://flybase.org/download/sequence/batch/?ids=FBgn0000490,FBgn0013765&type=FBpp
- http://flybase.org/download/sequence/batch/?ids=FBgn0000490,FBgn0013765&type=CDS&output=file
Links to Bulk Region Mode
For Bulk Region mode, you can pass in a space (%20) or semi-colon (%3b) delimited list of genomic sequence location strings, species, padding (additional flanking bases), and strand to populate the form with the specified values. Be sure to URL encode the space or semi-colon characters. Only genomic sequence location is required, all other parameters are optional.
- http://flybase.org/download/sequence/region/?regions=[Regions]&species=[Species]&padding=[Padding]&strand=[Strand]&output=[Output] (GET or POST)
- http://flybase.org/download/sequence/region/?regions=X:2790599..2796466
- http://flybase.org/download/sequence/region/?regions=X:2790599..2796466%3b4:69326..114270&padding=500&strand=minus
- http://flybase.org/download/sequence/region/?regions=X:2790599..2796466%3b4:69326..114270&padding=500&strand=minus&output=file